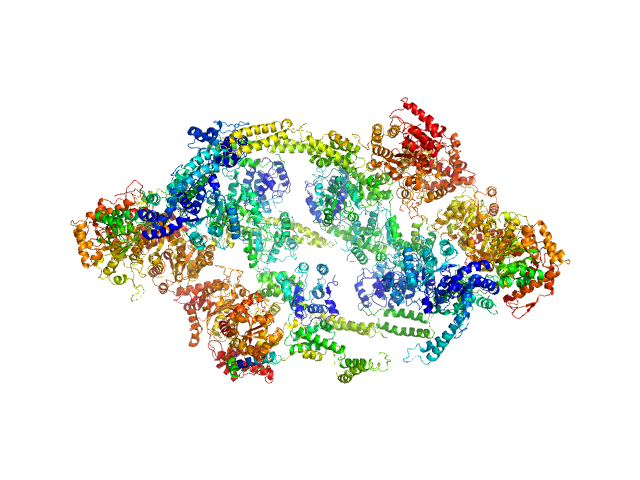
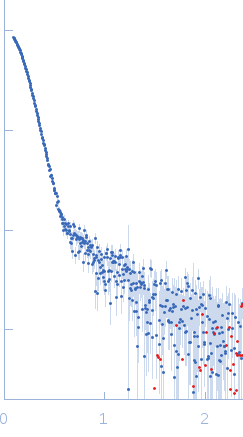
Linkage between ATP Consumption and Mechanical Unfolding during the Protein Processing Reactions of an AAA+ Degradation Machine: Cell

P2X7Rs decrease mortality, bacterial load, and inflammatory cytokines... | Download Scientific Diagram
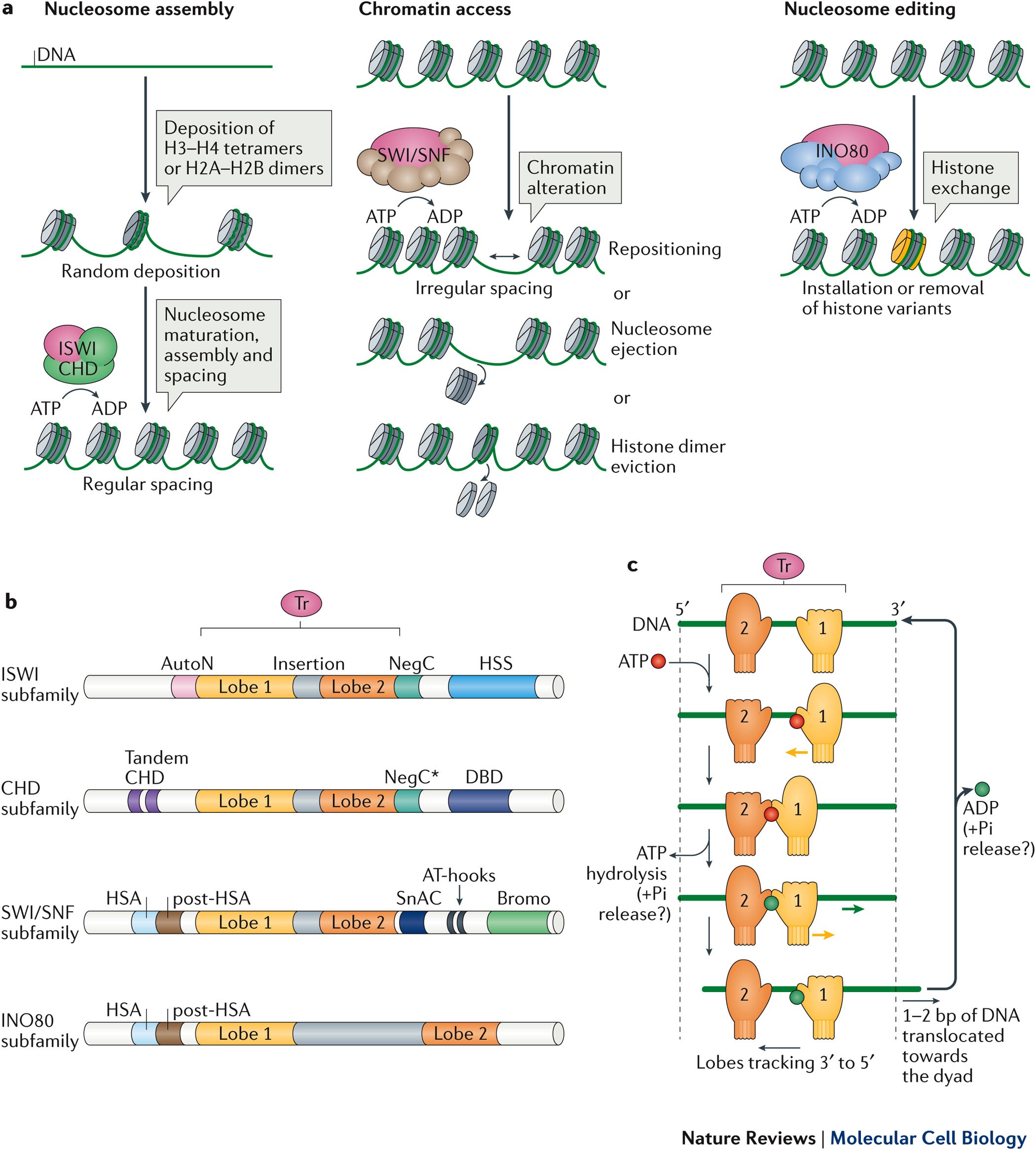
Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes | Nature Reviews Molecular Cell Biology
Registrants and notifiers to update their dossiers according to the provisions of the second ATP to CLP
IPI00003870 - CLPP Putative ATP-dependent Clp protease proteolytic subunit, mitochondrial precursor ENSG00000125656 - Detailed Information

RCSB PDB - 3FES: Crystal Structure of the ATP-dependent Clp Protease ClpC from Clostridium difficile
ClpX, an alternative subunit for the ATP-dependent Clp protease of Escherichia coli. Sequence and in vivo activities.